Parameter estimation is the process of determining model parameters using a dataset. This dataset can come from time course experiments, steady-state experiments, or a combination of both. COPASI is able to import a dataset, which may span across multiple files and contain several experiments.
Once you have loaded your dataset, COPASI fits one or more user-specified parameters to that data. The algorithms COPASI employs to estimate suitable parameter values are the same as those available in the optimization task. For more information on the various methods used, please refer to the methods section of this document.
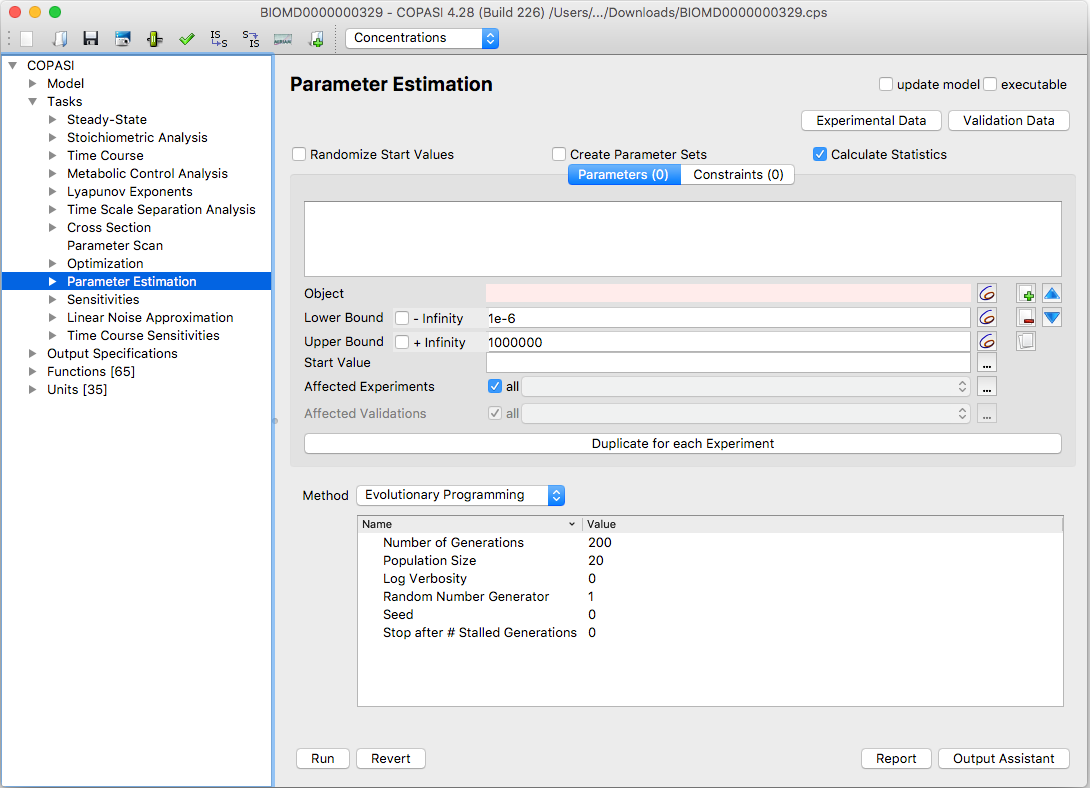 |
| Parameter Task Estimation Dialog |
If the Randomize Start Values checkbox is enabled, COPASI will generate a random start value within the defined bounds for each parameter you wish to estimate.
If the Create Parameter Sets checkbox is enabled, a new parameter set will
be saved for each defined experiment, containing the best solution found as well
as the original parameter values. Each parameter set is marked with the current
timestamp and can be found under
Model → Biochemical → Parameter Sets.
To open the parameter estimation dialog, select the Parameter Estimation branch under Multiple Tasks in the tree view on the left side of the user interface. You can define which parameters COPASI should fit—parameters are added in the same way as in the Optimization task. Click the button next to the Object field to open a selection dialog and choose the parameter you want to fit.
You can also specify upper and lower bounds for each parameter. COPASI will only fit parameters within these bounds. By default, the bounds are set to +Infinity (upper) and -Infinity (lower). To set your own bounds, uncheck the corresponding box and enter your desired value. Enter the lower bound in the field labeled “Lower Bound” and the upper bound in the field labeled “Upper Bound”.
For convenience, you may also enter bounds as percentages (e.g., -X% or +X%), which instructs COPASI to calculate the limit based on the start value. If needed, you can use the button next to the edit field to choose another model object as a bound for the parameter.
The start value is the initial parameter value that COPASI uses for fitting. By default, this is set to the current value from the model. You can override this by entering your own value or by using the options in the menu (opened with “…”) to reset to model values, randomize the value, or use the last estimated value. If the specified start value is outside the set bounds, COPASI will automatically adjust it to the nearest valid boundary during parameter estimation.
Additionally, you can restrict a parameter’s effect to subset(s) of your experiments. To do this, select the “…” next to Affected Experiments and choose the relevant experiments. For use cases where initial values need to be fitted separately for each experiment (such as different starting concentrations in time course data), the Duplicate for each Experiment button will create a copy of the current parameter for each specified experiment.